 |
 |
- Search
| Precis Future Med > Volume 5(2); 2021 > Article |
|
Abstract
Purpose
The purpose of this study was to calculate the cutoff value for amyloid β (Aβ) positron emission tomography (PET) positivity using the iterative outlier method and to evaluate its validity based on the concordance rate.
Methods
We performed the iterative outlier method on 373 cognitively unimpaired (CU) subjects and calculated the optimal cutoff value for Aβ positivity. The validation was performed using the independent dataset, comprising 83 subjects (27 CU, 27 amnestic mild cognitive impairment, and 29 Alzheimer’s dementia). We evaluated the validity of the Aβ cutoff value by calculating its concordance rate with the visual assessment and between two different Aβ tracers performed in the same subject.
Results
The concordance rate of Aβ cutoff values with the visual assessment ranged from 84.3% to 91.5%, depending on the reference regions. The concordance rate of the cutoff values between the two Aβ tracers ranged from 90.3% to 97.5%, all of which were higher than that of the visual assessment (86.7%).
Alzheimer’s disease (AD) is the most common cause of dementia in the elderly [1], and is characterized by the deposition of amyloid β (Aβ) in the brain. The amyloid cascade hypothesis, a well-known hypothetical model of AD, posits that accumulation of Aβ is the earliest pathogenic process leading to tau deposition, neurodegeneration, and clinical cognitive impairment [2]. Therefore, distinguishing brains free of Aβ from those with Aβ deposition is of utmost importance for the early diagnosis of AD [3].
Previous studies have presented a variety of approaches to define Aβ-positive cutoffs, including the iterative outlier method [7]. Until now, the validity of these cutoffs has often been evaluated with the standard-of-truth (SOT) based on either histological findings of brain autopsy or visual assessment of PET images. However, visual assessment is operator-dependent [8] and it is difficult to obtain postmortem data.
In this study, we calculated the cutoff value for Aβ positivity using the iterative outlier method and evaluated its validity based on the concordance rate. Two tracers were used to evaluate the concordance rate of Aβ positivity based on the cutoff value in the same subject.
For the iterative outlier method, we used Aβ PET data obtained from 373 cognitively unimpaired (CU) subjects aged over 50 years. CU subjects were defined as those with normal results on neurological examination and normal cognitive function, expected for their ages and education, on a standardized cognitive test (Seoul Neuropsychological Screening Battery) [9]. Of the 373 subjects, 202 and 171 subjects underwent 18F-florbetaben (FBB) and 18F-flutemetamol (FMM) Aβ PET, respectively.
For validation of the Aβ-positive cutoff value, we used the head-to-head comparison data, comprising 83 subjects (27 CU, 27 amnestic mild cognitive impairment, and 29 AD), in whom both FBB and FMM Aβ PET were performed. A detailed description is provided in our previous study [10].
Standardized three-dimensional T1 turbo field echo images were acquired from all subjects at the Samsung Medical Center using the same 3.0-T magnetic resonance imaging (MRI) scanner (Philips Achieva, Philips Healthcare, Andover, MA, USA). The following parameters were used: sagittal slice thickness of 1.0 mm, over contiguous slices with 50% overlap, no gap, repetition time of 9.9 ms, echo time of 4.6 ms, flip angle of 8°, and matrix size of 240 × 240 pixels, reconstructed to 480× 480 over a field of view of 240 mm.
PET images were acquired using a Discovery STe PET/computed tomography (CT) scanner (GE Medical Systems, Milwaukee, WI, USA) in three-dimensional scanning mode that examined 47 slices of 3.3-mm thickness spanning the entire brain. CT images were acquired using a 16-slice helical CT (140 KeV, 80 mA; 3.75-mm section width) for attenuation correction. According to protocols proposed by the ligand manufacturers, a 20-minute emission PET scan with dynamic mode (consisting of 4× 5 minutes frames) was performed 90 minutes after injection of a mean dose of 311.5 MBq FBB and 185 MBq FMM, respectively. Three-dimensional PET images were reconstructed in a 128× 128× 48 matrix with a voxel size of 2× 2× 3.27 mm using the ordered-subsets expectation maximization algorithm (FBB iterations= 4 and subset= 20; FMM iterations= 4 and subset= 20). Aβ positivity was determined based on visual assessments performed by three experienced raters (two nuclear medicine doctors and one neurologist) [11-15].
To calculate the SUVR, we performed the following processes. First, we co-registered the PET image to the T1-MRI images. Subsequently, we normalized the native PET image to the Montreal Neurological Institute-152 template using the transformation matrix, calculated during the segmentation of T1-MRI. After normalization, the brain was divided into 116 grey matter regions using the automated anatomical labeling (AAL) atlas [16]. We considered three reference regions (whole cerebellum [WC], cerebellar gray matter [CG], and pons) to calculate the SUVR, the regional masks of which were obtained from the GAAIN website (http://www.GAAIN.org) [17]. We calculated the global cortical SUVR as the volume-weighted mean of 56 cortical regions in the AAL template [18]. This pre-processing was performed using SPM8 through Matlab 2014b (Mathworks, Natick, MA, USA).
We used the iterative outlier method and generated an upper- and lower-bound SUVR. During the iteration, subjects with greater than the upper inner-bound (3rd quartile+ 1.5 interquartile range [IQR] or less than the lower inner-bound [1st quartile–1.5 IQR]) values were removed from the dataset. This process was repeated until all outliers were removed. In the final dataset, a cutoff value was determined by adding 2.5% of the maximum SUVR in itself [19].
We performed two analyses using the independent dataset to validate the Aβ cutoff value. We first compared the cutoff value-derived Aβ positivity with the visually determined Aβ positivity and calculated the concordance rate. Next, we evaluated whether the cutoff value-derived Aβ positivity of one subject was concordant between the two Aβ tracers (FBB and FMM). We assumed that the optimal cutoff value would distinguish subjects with positive Aβ deposition regardless of the tracer type. The statistical significance of the concordance rate was evaluated using the McNemar’s chi-square test, with a P-value of less than 0.05 indicating non-agreement between the two measures. All statistical analyses were completed using R version 3.5.3 (http://www.r-project.org/). This is a retrospective study using the data from the preceding study [10], which was approved by the Institutional Review Board of Samsung Medical Center (2015-04-091) with written informed consent of the participants at the time of enrollment.
Table 1 shows the demographics of the development and validation datasets. In the development dataset, subjects who underwent FBB were older than those who underwent FMM. In the validation dataset, although two different Aβ PETs were performed in the same subject, the frequency of A β positivity differed between the two tracers.
Using the iterative outlier method, we calculated the optimal cutoff value for Aβ PET positivity according to the Aβ tracers and reference regions (Fig. 1). In both FBB and FMM, the cutoff value was higher when the CG was used as the reference region.
To validate the cutoff value derived from the iterative outlier method, we applied the cutoff value for Aβ positivity in the independent dataset and compared it with that of the visual assessment (Table 2). The concordance rate ranged from 84.3% to 91.5%, and the highest rate was observed when the WC was used as the reference for both FBB and FMM. All P-values were greater than 0.05, indicating that the two measures (the cutoff value and the visual assessment) were in accordance.
We evaluated whether the cutoff value-derived Aβ positivity was consistent between the two tracers. The concordance rate ranged from 90.3% to 97.5%, all of which were higher than that of the visual assessment (86.7%).
The concordance rate of visual assessment between FBB and FMM was 86.7% (72/83) (Table 3, Fig. 2). There were 11 subjects who showed discordant results (nine subjects [FBB negative/FMM positive] and two subjects [FBB positive/FMM negative]). The concordance rates of the cutoff values were 90.3% (75/83), 97.5% (81/83), and 97.5% (81/83) for each reference region (CG, WC, and pons, respectively) (Table 3). The majority of the subjects who showed discordant results in the visual assessment showed concordant results when the cutoff value was used (9/11 for the CG [Fig. 2A], 11/11 for the WC [Fig. 2B], and 10/11 for the pons [Fig. 2C]).
In this study, we identified the optimal SUVR cutoff for Aβ positivity using the iterative outlier method and evaluated its validity based on the concordance rate.
We found that the cutoff optimization methods were comparable to visual assessment. Detecting individuals with Aβ deposition is important, as it is valuable for the clinical diagnosis of AD and in clinical trials aiming to reduce the burden of Aβ in the brain [4]. To date, in the clinical field, Aβ positivity of PET is generally determined by experienced clinicians based on visual rating. However, visual rating can be operator- and Aβ tracer-dependent, thus are prone to intra- and inter-rater discrepancies [8]. Interestingly, we found that the iterative outlier method-derived cutoff value had a higher concordance rate of Aβ positivity between the two tracers than that of the visual assessment. Although its superiority to visual assessment should be tested statistically, this finding suggests that the iterative outlier method-derived cutoff value could be an accurate measure of Aβ positivity in FBB and FMM PET.
Although the iterative outlier method may have an advantage over visual assessment, there are other approaches to define Aβ-positive cutoff values, such as receiver operating characteristic (ROC) analyses [20] and clustering methods [21]. The important difference in the approaches is that the iterative outlier method requires an annotated group of cognitively healthy subjects [7], whereas the ROC method requires both diseased and healthy groups [20], and the cluster analysis requires mixed groups that do not require annotations [21].
We further observed that cutoff optimization methods differed according to the reference regions and types of Aβ tracers. This discrepancy precluded the generalization of findings from one Aβ tracer to another [22]. Therefore, studies based on Aβ PET data should describe how the data is processed along with a description of the reference regions and the type of tracer.
The strengths of this study are that we used a data-driven approach to identify cutoff values and used a unique cohort (head-to-head comparison data) for the validation, where two different types of Aβ PET were performed in the same subject. However, this study has several limitations. First, we did not perform a statistical test to compare newly derived cutoff values with results of the visual assessment. This can be done by the bootstrapping method; however, the small sample size of our study precluded this analysis. Second, the iterative outlier method depends on the dataset used. Although we tried to obtain a homogenous dataset by thorough examinations, it is possible that the cutoff value will change if more subjects are included. Therefore, our findings should be interpreted with caution and replicated using a larger dataset. Lastly, ideally, histopathological confirmation of Aβ deposition in the brain should be the SOT. Due to the scarcity of post-mortem data, we considered visual assessment of Aβ PET as the SOT in this study.
In conclusion, we demonstrated that the iterative outlier method could identify the cutoff value for Aβ positivity in both FBB and FMM PET.
CONFLICTS OF INTEREST
No potential conflict of interest relevant to this article was reported. Sang Won Seo has been editorial board of P&FM since December 2017. He was not involved in the review process of this article.
Notes
AUTHOR CONTRIBUTIONS
Conception or design: HRK, YSC, SWS.
Acquisition, analysis, or interpretation of data: HRK, YSC, SHM, HJK, HJ, DLN, SP, SWS.
Drafting the work or revision: HRK, YSC, SWS.
Final approval of the manuscript: HRK, YSC, SWS.
Fig. 1.
Results of the iterative outlier method for (A, B, C) 18F-florbetaben (FBB) and (D, E, F) 18F-flutemetamol (FMM). Red lines indicate cutoff value derived using the iterative outlier method. SUVR, standard uptake value ratio; CG, cerebellar gray matter; WC, whole cerebellum.

Fig. 2.
Scatter plot of 18F-florbetaben (FBB) and 18F-flutemetamol (FMM) standard uptake value ratio (SUVR) with (A) cerebellar gray matter, (B) whole cerebellum, and (C) pons as the reference regions. Dotted lines indicate the cutoff value.
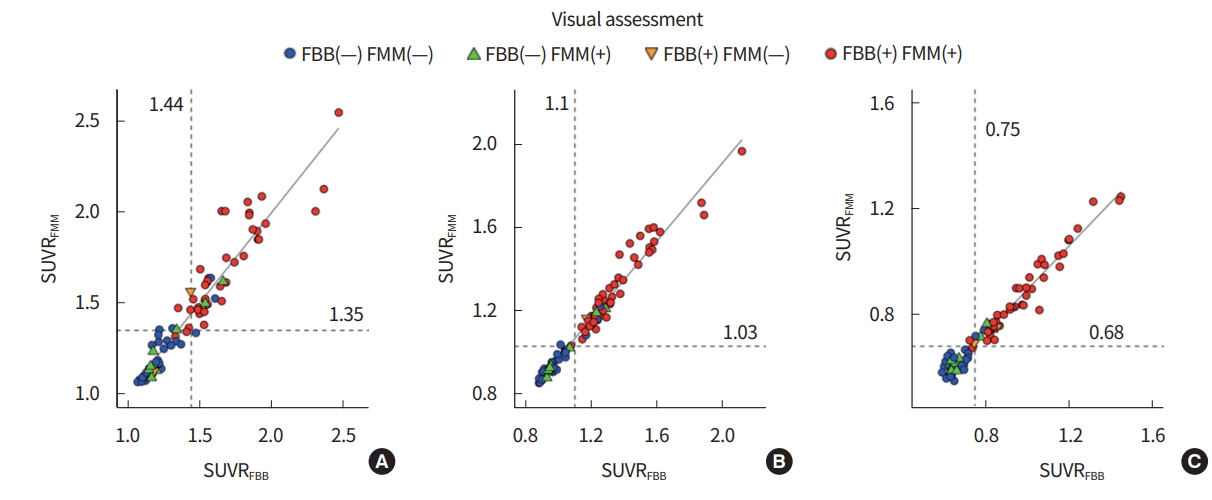
Table 1.
Demographics of the development and validation datasets
| Demographic |
Development dataset |
Validation dataset |
||
|---|---|---|---|---|
| FBB (n=202) | FMM (n=171) | FBB (n=83) | FMM (n=83) | |
| Age (yr) | 71.4±7.2 | 68.7±7.4 | 71.2±7.7 | |
| Women | 118 (58.4) | 109 (63.7) | 50 (60.2) | |
| Education (yr) | 12.6±4.4 | 11.6±5.0 | 9.8±4.8 | |
| MMSE, score | 28.3±1.5 | 28.0±2.1 | 25.5±5.0 | |
| Diagnosis (CU/aMCI/AD) | 202/0/0 | 171/0/0 | 27/27/29 | |
| Aβ positivitya) | 32 (15.8) | 27 (15.8) | 45 (54.2) | 52 (62.7) |
Table 2.
The concordance rate of the cutoff value with visual assessment
| Aβ tracer | Reference region | Concordance rate | P-valuea) |
|---|---|---|---|
| FBB | CG | 0.855 | 0.773 |
| WC | 0.915 | 0.450 | |
| Pons | 0.903 | 0.724 | |
| FMM | CG | 0.843 | 0.579 |
| WC | 0.855 | 0.773 | |
| Pons | 0.843 | 0.579 |
Table 3.
The concordance rate of the SUVR cutoff between the two Aβ tracers
| Methods | Concordance rate | P-valuea) |
|---|---|---|
| SUVR cutoff | ||
| CG | 0.903 | 0.077 |
| WC | 0.975 | 0.479 |
| Pons | 0.975 | 0.479 |
| Visual assessment | 0.867 | 0.070 |
REFERENCES
1. Alzheimer’s Association. 2018 Alzheimer’s disease facts and figures. Alzheimers Dement 2018;14:367–429.

2. Jack CR Jr, Knopman DS, Jagust WJ, Petersen RC, Weiner MW, Aisen PS, et al. Tracking pathophysiological processes in Alzheimer’s disease: an updated hypothetical model of dynamic biomarkers. Lancet Neurol 2013;12:207–16.



3. Sperling RA, Aisen PS, Beckett LA, Bennett DA, Craft S, Fagan AM, et al. Toward defining the preclinical stages of Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement 2011;7:280–92.



4. Blennow K, Mattsson N, Scholl M, Hansson O, Zetterberg H. Amyloid biomarkers in Alzheimer’s disease. Trends Pharmacol Sci 2015;36:297–309.


5. McNamee RL, Yee SH, Price JC, Klunk WE, Rosario B, Weissfeld L, et al. Consideration of optimal time window for Pittsburgh compound B PET summed uptake measurements. J Nucl Med 2009;50:348–55.


6. Logan J, Fowler JS, Volkow ND, Wang GJ, Ding YS, Alexoff DL. Distribution volume ratios without blood sampling from graphical analysis of PET data. J Cereb Blood Flow Metab 1996;16:834–40.


7. Aizenstein HJ, Nebes RD, Saxton JA, Price JC, Mathis CA, Tsopelas ND, et al. Frequent amyloid deposition without significant cognitive impairment among the elderly. Arch Neurol 2008;65:1509–17.



8. Haller S, Montandon ML, Lilja J, Rodriguez C, Garibotto V, Herrmann FR, et al. PET amyloid in normal aging: direct comparison of visual and automatic processing methods. Sci Rep 2020;10:16665.



9. Kang Y, Na D, Hahn S. Seoul neuropsychological screening battery. Incheon (KR): Human brain research & consulting Co.; 2003.
10. Cho SH, Choe YS, Kim YJ, Kim HJ, Jang H, Kim Y, et al. Head-to-head comparison of F-florbetaben and Fflutemetamol in the cortical and striatal regions. J Alzheimers Dis 2020;76:281–90.


11. Clark CM, Schneider JA, Bedell BJ, Beach TG, Bilker WB, Mintun MA, et al. Use of florbetapir-PET for imaging beta-amyloid pathology. JAMA 2011;305:275–83.



12. Farrar G. Regional visual read inspection of [F] flutemetamol brain images from end-of-life and amnestic MCI subjects. J Nucl Med 2017;58(Suppl 1):1250.
13. Johnson KA, Sperling RA, Gidicsin CM, Carmasin JS, Maye JE, Coleman RE, et al. Florbetapir (F18-AV-45) PET to assess amyloid burden in Alzheimer’s disease dementia, mild cognitive impairment, and normal aging. Alzheimers Dement 2013;9(5 Suppl):S72–83.



14. Kim HJ, Cho H, Werring DJ, Jang YK, Kim YJ, Lee JS, et al. F-AV-1451 PET imaging in three patients with probable cerebral amyloid angiopathy. J Alzheimers Dis 2017;57:711–6.

15. Martinez G, Flicker L, Vernooij RW, Fuentes Padilla P, Zamora J, Roque i Figuls M, et al. F-18 PET ligands for the early diagnosis of Alzheimer’s disease dementia and other dementias in people with mild cognitive impairment (MCI). Cochrane Database Syst Rev 2016 (5):CD012216.
16. Tzourio-Mazoyer N, Landeau B, Papathanassiou D, Crivello F, Etard O, Delcroix N, et al. Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. Neuroimage 2002;15:273–89.


17. Klunk WE, Koeppe RA, Price JC, Benzinger TL, Devous MD Sr, Jagust WJ, et al. The Centiloid Project: standardizing quantitative amyloid plaque estimation by PET. Alzheimers Dement 2015;11:1–15.


18. Park JH, Seo SW, Kim C, Kim SH, Kim GH, Kim ST, et al. Effects of cerebrovascular disease and amyloid beta burden on cognition in subjects with subcortical vascular cognitive impairment. Neurobiol Aging 2014;35:254–60.


19. Mormino EC, Brandel MG, Madison CM, Rabinovici GD, Marks S, Baker SL, et al. Not quite PIB-positive, not quite PIB-negative: slight PIB elevations in elderly normal control subjects are biologically relevant. Neuroimage 2012;59:1152–60.


20. Devanand DP, Mikhno A, Pelton GH, Cuasay K, Pradhaban G, Dileep Kumar JS, et al. Pittsburgh compound B (11CPIB) and fluorodeoxyglucose (18 F-FDG) PET in patients with Alzheimer disease, mild cognitive impairment, and healthy controls. J Geriatr Psychiatry Neurol 2010;23:185–98.



21. Bourgeat P, Chetelat G, Villemagne VL, Fripp J, Raniga P, Pike K, et al. Beta-amyloid burden in the temporal neocortex is related to hippocampal atrophy in elderly subjects without dementia. Neurology 2010;74:121–7.


22. De Santi S, Catafau A, Seibyl J, Bullich S. Robustness of F-Florbetaben SUVR cutoff quantification across reference regions and standards of truth. J Nucl Med 2016;57(Suppl 2):458.







